Introduction
Hippophae rhamnoides L commonly called Sea Buckthorn (SBT) is a valuable and unique natural sources gained attention worldwide not only for its medicinal properties but also for its nutritional properties. It is a multi-medicinal tree shrub of the genus Hippocea belongs to the family Elaeagnaceae. Hippophae rhamnoides L. is mostly found in the moderate geographic locations of the north hemisphere, categorized into six species and 12 subspecies in the plant taxonomy [1]. It is a deciduous and nitrogen-fixing plant of cold arid region, native to Asia and Europe. All parts of SBT are considered as a rich source of numerous bioactive constituents with excessive nutritional and medicinal properties [2-4]. Due to these properties, H. rhamnoides L. is domesticated around the globe [5,6]. It is a very important medicinal remedy which is considered to be a rich source of a large number of phytochemicals, nutrients and bio-active constituents [7,8]. The bio-active compounds include vitamins, carotenoids, phytosterols, poly unsaturated fatty acids, organic acids, mineral components and essential amino acids [7-10]. SBT has been used in Europe and Asia for pharmaceuticals, foods and therapeutic purpose for centuries. SBT leaves, berries and seeds are well known for their medicinal properties [11-14]. The known medicinal properties of SBT include immune-modulatory, anti-stress, radio protective, antioxidant, tissue regeneration, anti-atherogenic, hepato-protective property and improving functions of blood circulation and digestive system [4,15-26]. It is interesting to note that the aqueous extract of SBT has recently been reported to possess hypoglycemic activity [27]. SBT contains a lot of different bioactive constituents with multiple properties which have ability to prevent postprandial hyperglycemia [28]. Flavonoids from fruits and seeds of SBT can cause hypolipidemia and hypoglycemia [29]. The leaves of methanolic extracts of SBT contain compounds showing alpha-glucosidase inhibition activity which may be used in diabetes because it has the ability to control inhibition activity [30].
Methodology
General instrumentation
UV spectra were obtained using a Hitachi-U-3200 spectrophotometer. IR spectra were recorded on a Jasco A-302 spectrophotometer. 1D- and 2D NMR spectra were obtained on AM-500, Bruker spectrometers in CD3OD and CDCl3, using tetramethylsilane (SiMe4) as internal standard. Mass spectra (EI-MS and HR-EIMS) were analyzed on a Mass AB SCIEX spectrometer QSTAR xl. Column chromatography was carried out by using silica gel 60 (Merck, 70–230 mesh).TLC was carried out on silica gel 60 PF254 (Merck), with detection by UV at 254 nm and 366 nm.
Plant material
The Sea buckthorn berries (8.5kg) were collected randomly in August-September from Gilgit, north of Pakistan in September 2015. The plant was identified by a botanist from Department of Botany, University of Karachi. A voucher specimen (No.G.H.No. 84346) was been deposited in the Herbarium, Department of Botany, University of Karachi.
Extraction and isolation
The berries were dried at room temperature and soaked in methanol for 72h (3times). The extract solution was filtered and concentrated under vacuum. The extract was suspended in water (0.5L) and partitioned with ethyl acetate to give the ethyl acetate-soluble part (111.0g). The ethyl acetate fraction was then fractionated with 4% Na2CO3 and 30% HCl to obtain acidic and basic fractions. The basic fraction was dried over Na2SO4 washed with water and evaporated the solvents under reduced pressure and obtained a neutral fraction. Both the acidic and neutral fractions were partioned with hexane to get hexane insoluble and soluble fractions.
The ethyl acetate acidic hexane insoluble part (10g) was subjected to a silica gel column chromatography, using a gradient solvent system of Pet. ether EtOAc CH2Cl2 MeOH with increasing polarity to give 20 fractions which were combined on the basis of TLC and obtained eight sub-fractions (HRF 1-8). The fraction 2 (1.02g) was fractionated into four sub-fractions through normal phase CC. Compound 1 was purified from the fraction HRF-2D using hexaneethyl acetate (55 vv) as solvent system. Fraction 4 (1.48g) was further subjected to column chromatography using Pet. etherEtOAc as solvent system (8.51.5vv) and obtained compound 2.Compound 3 was purified from fraction 4 using CH2Cl2 MeOH (91 vv) as solvent system through normal phase CC (Figure 1).
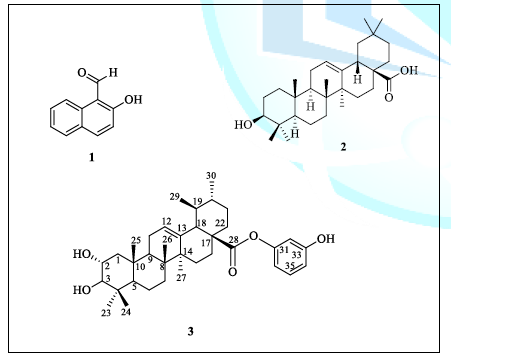
Figure 1: Structure of compounds 1, 2 and 3.
Sulforhodamine B assay
The growth inhibitory activities of compounds 1, 2, and 3 were evaluated against human non-small cell lung (NCI-H460) and breast (MCF-7) cancer cell lines by using sulforhodamine-B assay for the determination of IC50 values. The concentration of the test agents that inhibits 50% of the cell growth was referred as IC50. For this, the cells (10000cells100µL) from respective cell lines were incubated in 96 well plates for 24h at 37ºC in 5% humidified CO2 incubator. The stock solutions of compounds 1, 2, and 3 (20 mM) in DMSO, and doxorubicin (1mM) in distilled water were prepared. A range of dilutions for 1 (1, 10, 25, 50 and100µM), 2 and 3 (1, 2.5, 5, 7.5 and 10 µM) were added (100µL) in respective wells. The pre-determined IC50 value of doxorubicin (5µM) was used as positive control (data for range of dilutions are not shown). After completion of 48h, ice cold TCA (50µL, 50%) was added and left at room temperature for 30min. The TCA was washed out with distilled water. After drying, SRB solution (100µL, 0.4% wtvol in 1% acetic acid) was added to each well and unbound stain was washed out after 30min with1% acetic acid. Tris-base (100 µL, 10 mM, pH 10.2) was added to each well to solubilize protein bound stain. The absorbance for test agents along with appropriate blanks (both test agent and control) was recorded at 545nm using a microplate reader (Synergy). This absorbance was used to calculate respective percent growth inhibitions of the test agents. The dose response curve graph was plotted between growth inhibitions (y-axis) vs concentrations of the respective test agents (x-axis) to determine IC50 value.
DNA docking studies
To perform the DOCK calculations that predict the best orientations of the ligand in the binding site of the receptor we require preparing the receptor and ligand as inputs. UCSF Chimera is a visualizing tool used here to prepare the receptor and ligand with its Dock Prep tool option [31]. The structure of DNA PDB ID 1d29 was selected as receptor and downloaded from Protein Data Bank (PDB) [32]. The three compounds are taken as ligands to check if they are making any interactions with the DNA moiety. Hydrogen atoms and partial charges were added to both ligand and receptor. AM1-BCC charges were added to receptor, gasteiger charges were added to ligand and files were saved in mol2 format as program DOCK read file in Mol2 format. Molecular surface of receptor was prepared by DMS program. The DOCK accessory program sphgen was used to generate the spheres with a probe radius of 1.4 angstrom. It generated 12 clusters and the cluster 1 with maximum number of spheres i.e. 38 was selected for construction of box with the help of show box DOCK accessory program. The maximum 38 spheres in cluster 1 were retained for docking (Figure 2).
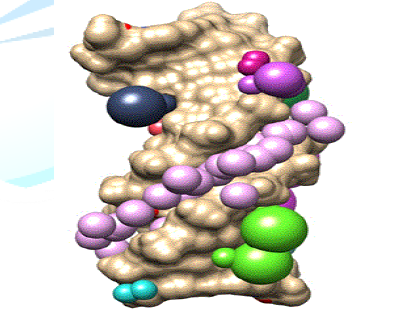
Figure 2: Surface diagram of DNA (PDB ID: 1d29) representing the cluster 1 with maximum numbers of violet colored spheres.
Following
the construction of box, the DOCK accessory program grid was used to
pre-compute the energy interaction points with grid spacing of 0.3 angstrom
resolution within the box [33]. The number of grid points on x, y, and z axis
was 109, 85, and 108 Å, respectively with box dimension 32.375, 25.076, 32.008Å
and center on 12.814, 23.837, and 4.833 Å (Figure
3). Once the grid box was set we performed the docking by treating the
ligand as flexible. All the parameters were set as default value given in
standard protocols file. The maximum number of orientation was set as 1000 and
maximum number of conformations was kept 10 for each compound.
Electrophoretic mobility of plasmid pBR322 DNA
Electrophoretic
mobility of pBR322
plasmid DNA was performed to assess the direct interaction of compounds with
DNA. Briefly, agarose gel (1%) was prepared in 1x TAE buffer (70mL) by heating
for 1min followed by its casting in gel tray at room conditions. pBR322 DNA (500ng, 5µLwell) was mixed
either with PBS (control) or with compounds 1, 2, 3 (2.5µM), and doxorubicin (250 nM) and placed at 37ºC for
30min. Loading buffer (3µL, bromophenol blue, xylene cyanol and glycerol in a
ratio of 11120) was applied to the above reaction mixture. This sample was
loaded in the wells of the agarose gel and electrophoresis was performed in TAE
buffer at 70 V for 1.5h. The gel was dipped in ethidium bromide solution
(5μgµL) for 20min and washed with tap water. The mobility pattern
of the circular DNA was observed under UV light and
photographed.
Statistical analysis
Duncan’s multiple range test using SPSS 17 program. The data was analyzed
by using one-way
ANOVA with p<0.05 level was considered as significant
followed by.
Results and Discussion
Compound 1
was purified as white powder having the molecular formula C11H8O2
([M+] mz 172.05). The IR spectrum showed absorption peaks at 3486cm-1.The
1H-NMR shows four doublets of the aromatic proton at δ 8.32 (d, J = 8.5 Hz,
1H), δ 7.96 (d, J = 7.9 Hz, 1H), δ 7.78 (d, J = 8.5 Hz, 1H), δ 7.59 (m, 1H), δ
7.41 (m, 1H) and δ 7.12 (d, J = 9 Hz, 1H). Two singlets at δ 10.792 and 13.14
were appeared for aldehyde proton and hydroxyl proton resp.Compound 2 was isolated as white amorphous
powder. The molecular ion [M+] peak was appeared at mz 456.4 in
EI-MS spectrum matching to C30H48O3. Other
characteristic peaks were appeared at mz 248, 203 and 133. In ultra-violet
spectrum, an absorption band was appeared at 213 nm. The infra-red spectrum of
compound 2 shows absorption peak in the area of 2940.9cm-1 for
symmetric vibrations of CH2cm-1 group and at 1696.1cm-1
appears a characteristic band of carbonyl group (C=O). A broad peak was also
appeared in the area of 3446.7 cm-1 for hydroxyl group. The 1H-NMR
spectrum of compound 2 revealed
seven singlets of tertiary methyl protons at δ 0.66, 0.67, 0.79, 0.79, 0.81,
0.86 and 1.03. A doublet of doublet of C-18 proton at δ 2.71 and a triplet of
one vinyl proton of C-12 at δ 3.09 indicates an olea-12-ene skeleton. One broad
singlet peak at δ 4.8 showed the signal of O-H group. The above mentioned
spectral data of compound 2 were in complete agreement with those reported in
literature for oleanolic acid [34].
Compound 3 was obtained as white
amorphous solid and revealed to have the molecular formula C36H52O5
([M]+ at mz 564.3814, calculated 564.3815) by HREI-MS. The fragment
ion peaks at mz 472.3 [M-C6H5O]+ showed the
loss of phenol moiety. Two fragment ion peaks at mz 454.4 [M-H2O]+
and at mz 437.3 [M-2H2O]+ indicate that compound 3 contained two hydroxyl groups.The UV
spectrum showed absorption band at 230 and 298 nm. In infra-red spectrum, a
very intensive absorption peak in the area of 2932 cm-1 for
symmetric vibrations of CH2 group and at 1699 cm-1
appears a characteristic band of carbonyl group (C=O). A broad peak was also
appeared in the area of 3289 cm-1 for hydroxyl group. The 1H
NMR spectra of compound 3 showed the
presence of five methyl signals at δ 0.75, 0.99, 1.05, 1.09, and 1.17 two
methyl doublets that appeared at δ 0.92 and 1.01. The 1H NMR spectra
of compound 3 also showed two
oxymethine protons resonating at δ 3.32 and 3.37 and a olefinic proton at δ
5.21. The
presence of five methyl singlets and two methyl doublets suggested that compound
3 belongs to ursane type
triterpenoid having two secondary hydroxyl groups and a tri substituted double
bond between C-12C-13. In aromatic range, two doublet of doublet appeared at δ
6.97 and 7.48. The spectrum also displayed a multiplet at δ 7.22 and a doublet
at δ 7.34. The 13C-NMR values
for all the protons and carbons were assigned on the basis of HMQC and HMBC
correlations and were given in Table 1.
The appearance of an ester carbonyl group resonating at δ 180.1 in the 13C
NMR spectral data of 3 suggested the
presence of an ester functional group and its location was identified at C-28
by the key COSY and HMBC correlations as shown in Figure 4.
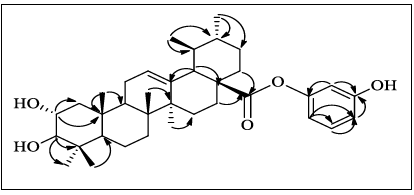
Figure 4: HMBC correlation of compound 3.
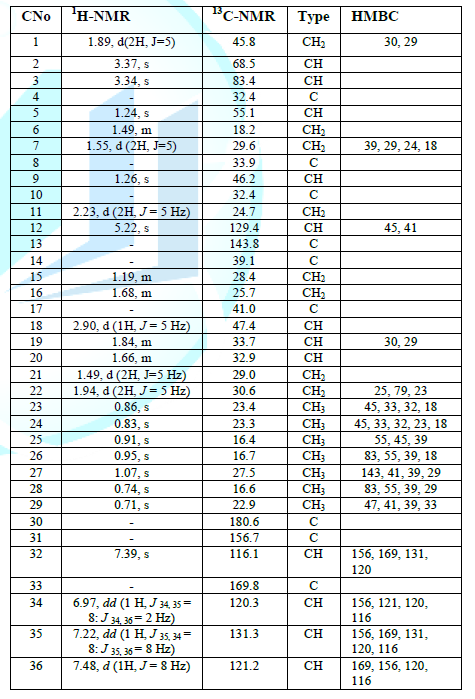
Table 1: NMR spectral data of compound 3 (1H-NMR 500 Hz, 13C 125 MHz).
A search in literature found that the spectral
characteristics of 3 were consistent
to the reported literature values of Corosolic acid except the aromatic
spectral data that confirmed by the key COSY and HMBC correlations as shown in Figure 3. The correlation between
protons and carbon of compound 3 are
summarized in table 1.
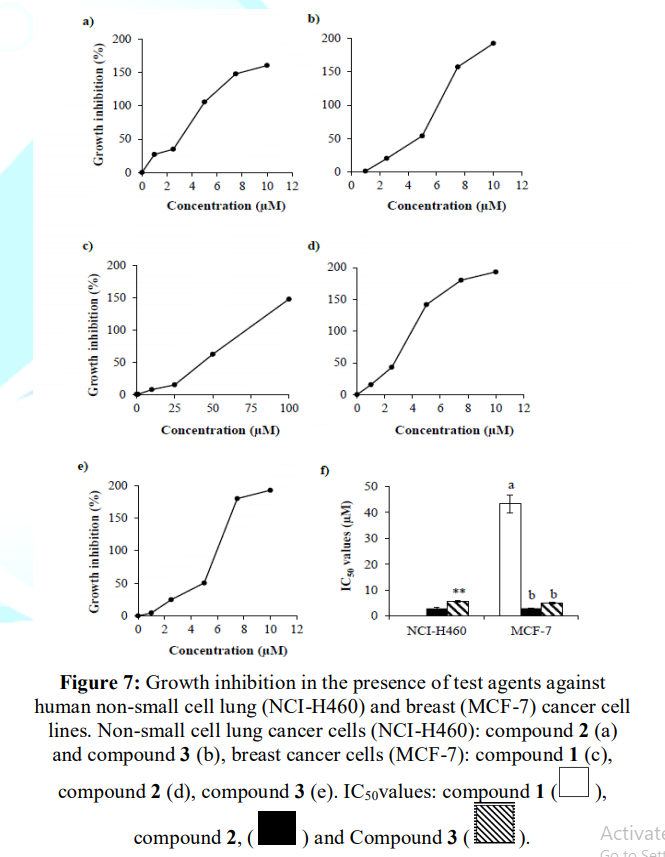
Figures 5 and 6 depicts compound 1 was not effective against non-small
cell lung
cancer cell line (NCI-H460) while it displayed growth
inhibition against breast cancer cells (MCF-7) with IC50 value of
~43 µM. In addition, compound 2 and 3 exhibited growth inhibition against
both non-small cell lung and breast cancer cells. In lung cancer cell line
(NCI-H460), HI-6 exhibited IC50 value of ~2.8 µM which was ~2x more
potent than that of HI-7. However, in case of breast cancer cells (MCF-7), both
were equipotent with IC50 value of ~3 µM (Figure
7).
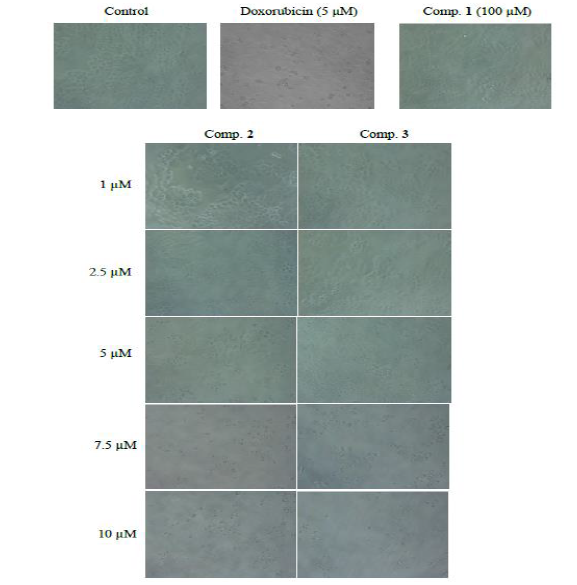
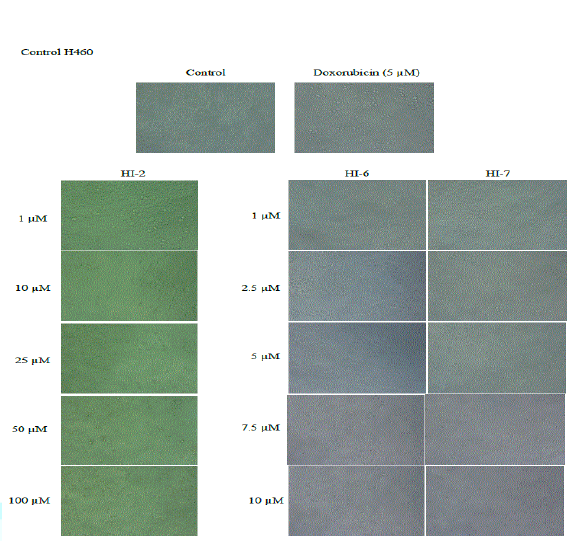


Figure 6: Growth inhibitory effects of comp. 1, 2 and 3against breast cancer (MCF-7) cell line.
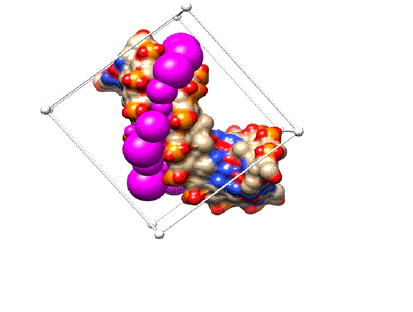
After successful docking runs, the best docked conformation for each compound was viewed in UCSF chimera by using View Dock tool option. All three compounds were docked in the minor groove of DNA model (Figure 8).
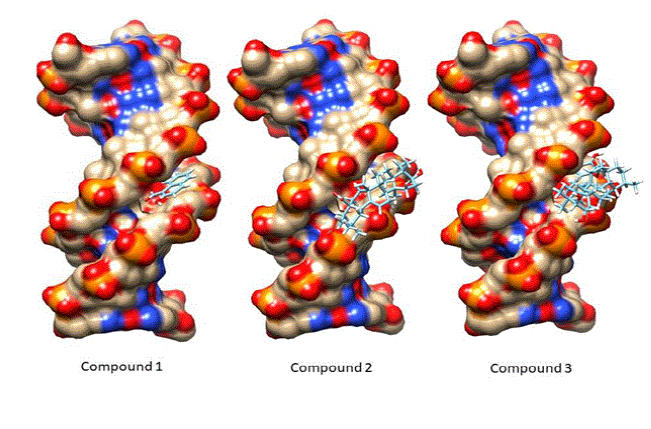
Figure 8: All three compounds docked in the minor groove of DNA.
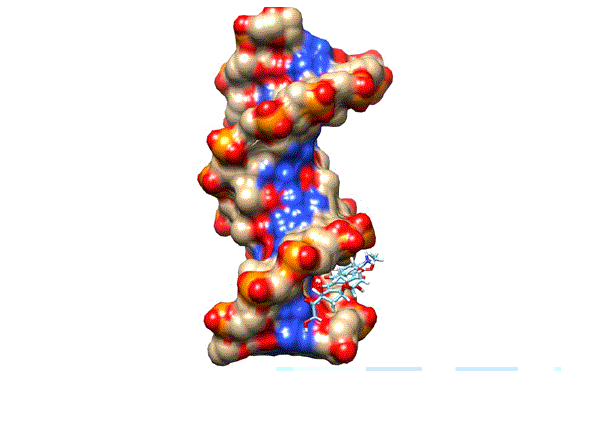
Figure 9: The docking of doxorubicin in the minor groove of DNA.
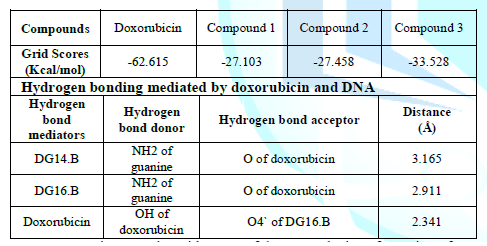
Structure analysis of each compound was done by Structural analysis tool. We did not find any hydrogen bond interactions between DNA and our top ranked conformations of three compounds. They formed weak interactions such as van dar waal and hydrophobic interactions with the DNA. We also docked the Doxorubicin by following the same protocols. It produces its anticancer activity through intercalation mode. The doxorubicin was also docked in the minor groove but at the different position (Figure 9). The docking of doxorubicin in minor grooves suggests that minor groove binding is the dominant pre-intercalation step (Lei, 2012 #17). The rigid scores of all compounds and doxorubicin are given in Table 2. The high negative value of doxorubicin indicates that it is a strong DNA binder than our compounds. The docking outcomes also correlate with the gel electrophoresis run where doxorubicin is showing high affinity as compared to our three compounds.
The energy score of compound 3 has a relatively high negative value than compounds 1 and 2. In case of doxorubicin while performing the structure analysis with the help of UCSF chimera, three hydrogen bonds were found. The purine bases guanine (DG14.B and DG16.B) of strand B are involved in mediating the hydrogen bonding shown in Figure 10. The details of the hydrogen bond mediators, donors, acceptor and distances of doxorubicin and DNA are given in table 2.
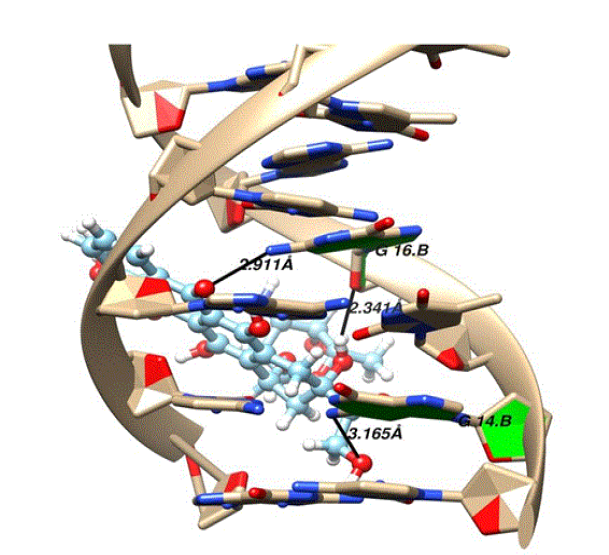
Figure 10: The three hydrogen bond interactions formed between doxorubicin and DNA.
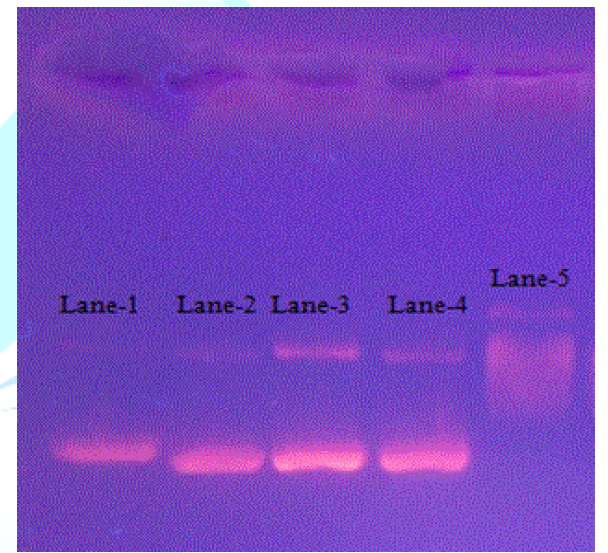
Figure 11 depicts that in control (lane-1), two bands of similar shape were observed and no change was observed in the presence of 1, 2, 3 at 2.5μM (lanes -2, -3 and -4, respectively). However, at 250nM, doxorubicin (lane-5) exhibited smear formation suggesting its interaction and damaging effect on DNA. From the DNA docking and electrophoretic mobility experiments, it is noted that anticancer activity of these three compounds is not related to DNA intercalation. However, weak interactions of these compounds in the minor groove of DNA suggest that these may modify the DNA function at the level of gene expression [35]. Therefore, further studies at transcriptional level should be performed to determine the mechanism of their anticancer activity, whereas the docking result of doxorubicin suggests clearly that it is a strong DNA binder rendering it as an anticancer agent.
Conclusions
It is concluded that the new compound 3 was isolated from the sea buck thorn. It displayed potent anticancer activity against lung cancer than that of known compounds 1 and 2 with no structural DNA damage. Their weak interactions with minor groove of DNA suggest that they may modify gene expression to induce anticancer activity. Lahore, Pakistan. The plant materials leaf, stems and seeds were properly cleaned and kept in room temperature.
References
- Bal LM, Meda V, Naik S, Satya S. Sea buckthorn berries: a potential source of valuable nutrients for nutraceuticals and cosmoceuticals (2011) Food Res Int 44:1718-1727. https://doi.org/10.1016/j.foodres.2011.03.002
- Malinowska P, Olas B. Sea buckthorn-valuable plant for health (2016) Kosmos 65:285-292.
- Shivapriya S, Ilango K, Dubey G. Evaluation of antioxidant and neuroprotective effect of Hippophae rhamnoides (L.) on oxidative stress induced cytotoxicity in human neural cell line IMR32 (2015) Saud J Biol Sci 22:645-650.https://doi.org/10.1016/j.sjbs.2015.04.011
- Upadhyay NK, Kumar R, Siddiqui M, Gupta A. Mechanism of wound-healing activity of Hippophae rhamnoides L. leaf extract in experimental burns (2011) Evidence-based complementary and alternative medicine. https://dx.doi.org/10.1093/ecam/nep189
- Li TS, Beveridge TH. Sea Buckthorn (Hippophae rhamnoides L.) Production and Utilization (2003) NRC Research Press, Canada.
- Rousi A. The genus Hippophae L. A taxonomic study Paper (1971) Annales Botanici Fennici 8:177-227.
- Beveridge T, Li TS, Oomah BD, Smith A. Sea buckthorn products: manufacture and composition (1999) J Agri Food Chem 47:3480-3488. https://doi.org/10.1021/jf981331m
- Zeb A. Chemical and nutritional constituents of sea buckthorn juice (2004) Pakistan J Nutrition 3:99-106. http://dx.doi.org/10.3923/pjn.2004.99.106
- Mäkinen KK, Söderllng E. A quantitative study of mannitol, sorbitol, xylitol, and xylose in wild berries and commercial fruits (1980) J Food Sci 45: 367-371. https://doi.org/10.1111/j.1365-2621.1980.tb02616.x
- Yang B, Kallio HP. Fatty acid composition of lipids in sea buckthorn (Hippophaë rhamnoides L.) berries of different origins (2001) J Agri Food chem 49:1939-1947. https://doi.org/10.1021/jf001059s
- Geetha S, Jayamurthy P, Pal K, Pandey S, Kumar R, et al. Hepatoprotective effects of sea buckthorn (Hippophae rhamnoides L.) against carbon tetrachloride induced liver injury in rats (2008) J Sci Food Agri 88:1592-1597. https://doi.org/10.1002/jsfa.3255
- Hsu YW, Tsai CF, Chen WK, Lu FJ. Protective effects of seabuckthorn (Hippophae rhamnoides L.) seed oil against carbon tetrachloride-induced hepatotoxicity in mice (2009) Food Chem Toxicol 47:2281-2288. https://doi.org/10.1016/j.fct.2009.06.015
- Sabir SM, Maqsood H, Hayat I, Khan M, Khaliq A. Elemental and nutritional analysis of sea buckthorn (Hippophae rhamnoides ssp. turkestanica) berries of Pakistani origin (2005) J Med Food 8:518-522. https://doi.org/10.1089/jmf.2005.8.518
- Zeb A. Anticarcinogenic potential of lipids from Hippophae: evidence from the recent literature (2006) Asian Pacific J Cancer Prevent 7: 32.
- Basu M, Prasad R, Jayamurthy P, Pal K, Arumughan C, et al. Anti-atherogenic effects of seabuck thorn (Hippophaea rhamnoides) seed oil (2007) Phytomedicine 14:770-777. https://doi.org/10.1016/j.phymed.2007.03.018
- Chawla R, Arora R, Singh S, Sagar RK, Sharma RK, et al. Radioprotective and antioxidant activity of fractionated extracts of berries of Hippophae rhamnoides (2007) J Med Food 10: 101-109. https://doi.org/10.1089/jmf.2006.007
- Gao ZL, Gu XH, Cheng FT, Jiang FH. Effect of sea buckthorn on liver fibrosis: a clinical study (2003) World J Gastroenterology 9:1615-1617. https://dx.doi.org/10.3748%2Fwjg.v9.i7.1615
- Geetha S, Ram MS, Singh V, Ilavazhagan G, Sawhney R. Anti-oxidant and immune-modulatory properties of seabuck thorn (Hippophae rhamnoides) - an in vitro study (2002) J Ethno pharma 79:373-378. https://doi.org/10.1016/S0378-8741(01)00406-8
- Geetha S, Ram MS, Singh V, Ilavazhagan G, Sawhney R. Effect of seabuck thorn on sodium nitroprusside-induced cytotoxicity in murine macrophages (2002) Biomed pharmaco therapy 56:463-467. https://doi.org/10.1016/S0753-3322(02)00290-1
- Goel H, Prasad J, Singh S, Sagar R, Kumar IP, et al. Radioprotection by a herbal preparation of Hippophae rhamnoides, RH-3, against whole body lethal irradiation in mice (2002) Phyto med 9:15-25. https://doi.org/10.1078/0944-7113-00077
- Gupta A, Kumar R, Pal K, Banerjee PK, Sawhney RC. A preclinical study of the effects of seabuck thorn (Hippophae rhamnoides L.) leaf extract on cutaneous wound healing in albino rats (2005) The Int J Low Extrem Wounds 4:88-92. https://doi.org/10.1177/1534734605277401
- Saggu S, Divekar HM, Sawhney RC, GuptaV, Banerjee PK, et al. Adaptogenic and toxicity evaluation of Sea buckthorn (Hippophae rhamnoides) leaf extract: a dose dependent study (2006) Toxicol Lett 164:S196.https://doi.org/10.1016/j.toxlet.2006.07.066
- Süleyman H, Demirezer L, Büyükokuroglu M, Akcay M, Gepdiremen A, et al. Antiulcerogenic effect of Hippophae rhamnoides L (2001) Phyto Ther Research 15:625-627. https://doi.org/10.1002/ptr.831
- Upadhyay N, Kumar R, Mandotra S, Meena R, Siddiqui M,et al. Safety and healing efficacy of Sea buckthorn (Hippophae rhamnoides L.) seed oil on burn wounds in rats (2009) Food Chem Toxico 47:1146-1153. https://doi.org/10.1016/j.fct.2009.02.002
- Xing J, Yang B, Dong Y, Wang B, Wang J, et al. Effects of sea buckthorn (Hippophae rhamnoides L.) seed and pulp oils on experimental models of gastric ulcer in rats (2002) Fitoterapia 73: 644-650. https://doi.org/10.1016/S0367-326X(02)00221-6
- Yang B, Kallio H. Composition and physiological effects of sea buckthorn (Hippophae) lipids (2002) Trends Food Sci Tech 13:160-167. https://doi.org/10.1016/S0924-2244(02)00136-X
- Zhang W, Zhao J, Wang J, Pang X, Zhuang X, et al. Hypoglycemic effect of aqueous extract of seabuck thorn (Hippophae rhamnoides L.) seed residues in streptozotocin induced diabetic rats (2010) Phyto Ther Res 24:228-232. https://doi.org/10.1002/ptr.2917
- Kim YM, Wang MH, Rhee HI. A novel α-glucosidase inhibitor from pine barks (2004) Carbohydrate Res 339:715-717.
- Cao Q, QuW, Deng Y, Zhang Z, Niu W, et al. Effect of flavonoids from the seed and fruit residue of Hippophae rhamnoides L. on glycol-metabolism in mice (2003) J Chinese Medi Mat 26:735-737. https://doi.org/10.1016/j.carres.2003.11.005
- Bhardwaj P, Varshneya C, Kaistha K, Tandon T. In vitro evaluation of anti-diabetic and antioxidant activity of Seabuckthorn (Hippophae rhamnoides L.) leaves (2015) J Medi Plants Re 9:929-932. https://doi.org/10.5897/JMPR2015.5416
- Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, et al. UCSF Chimera-a visualization system for exploratory research and analysis (2004) J Comput Chem 25:1605-1612. https://doi.org/10.1002/jcc.20084
- Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, et al. International tables for crystallography volume F: crystallography of biological macromolecules (2006) Springer, USA.
- Allen WJ, Balius TE, Mukherjee S, Brozell SR, Moustakas DT, et al.Impact of new features and current docking performance (2015) J Comput Chem 36:1132-1156. https://doi.org/10.1002/jcc.23905
- Onoja E, Ndukwe I. Isolation of oleanolic acid from chloroform extracts of Borreria stachydea (2013) J Nat Prod Plant Resour 3:57-60.
- Baraldi PG, Bovero A, Fruttarolo F, Preti D, Tabrizi MA, et al. DNA minor groove binders as potential antitumor and antimicrobial agents (2004) Med Res Rev 24:475-528. https://doi.org/10.1002/med.20000
*Corresponding author:
Huma Aslam Bhatti, HEJ Research Institute of Chemistry International Center for Chemical and Biological Sciences, University of Karachi, Karachi, Pakistan, Tel: +92 21 99261779, Fax: +92 21 34819018-19, E-mail: huma_aslam31@hotmail.com
Citation:
Keywords
Anti-cancer, Corosolic acid, DNA, Hippophae rhamnoides, Pentacyclic triterpene


 PDF
PDF