Research Article :
Cannon L Mark, Kabat B,
Yogev R, Jantra L, Awan A, Le C,
White K and Vorachek A The interaction between the human
host micro biome and over the counter products has recently been investigated,
with surprising results. Some over the counter items may negatively affect the
health of the host, supporting the concept of the “hygiene hypothesis”, that
is, that disease may be actually caused by the lack of beneficial commensal
bacteria. Recent reports on the gluten metabolizing genus, Rothia, and a possible association with Celiac
Disease beg the question, what happened to the Rothia? In this study inhibitory factors, such as, Over The
Counter oral hygiene products and antagonistic bacteria were investigated and, in vitro, significantly inhibited the
gluten metabolizing bacteria, possibly affecting human digestion and
contributing to gluten sensitivity. The human body is host to trillions of microorganisms,
including bacteria, molds, yeasts, viruses and archaea. In addition, the
contribution of the microbiome to human health has become thoroughly
established with roles such as educating the immune response, resisting pathogens,
and digestion. As a result, the human microbiome project was designed to
ascertain the microbial composition of the entire human body. Meanwhile, the
oral microbiome has been extensively determined and reported in the literature.
The current reported microbiome of the oral cavity region contains 619 taxa,
derived from 13 phyla [1-4].
An additional 36,043 gene clones have been sequenced,
identifying an additional 434 unique oral taxa that (after further validation)
may be added to the database. Amongst the oral strains sequenced to date, two
important gluten metabolizing species,
Rothia mucilaginosa and Rothia aeria have been identified [4,5]. R.
mucilaginosa and R. aeria are
of the Rothia genus under the phyla Actinobacteria. R. aeria was named after its isolation from air in the Russian
space laboratory Mir and is an oral inhabitant [6,7]. R. mucilaginosa is primarily found in the oral
cavity but has been reported in the upper respiratory tract and also the
duodenum [8-11]. Interestingly, mucosal damage in celiac disease is mostly
found in this area of the gastro-intestinal system [12]. Oral micro-organisms
that in vitro degrade dietary
proteins may mean that they play an in
vivo role in food metabolism.
During mastication, ingested food is mixed with stimulated whole saliva and
oral micro-organisms. This process accelerates food digestion while the bolus
is still churning in the oral cavity [13]. For example, nitrate reducing bacteria have been described
as being indispensable in the production of nitric oxide which regulates blood
pressure and cardiovascular health and this further emphasize the importance of
the oral microbiome in systemic health [14-16]. A favorable and potential
source for gluten-degrading
enzymes would be the micro-organisms inhabiting the human gastro-intestinal
tract. It is well reported that bacteria residing in and on the human body
supply the host with numerous functions that are not encoded by the human
genome [17]. For instance, bacteria that colonize the large intestine ferment
starches that are resistant to mammalian digestive enzymes [18]. In addition, it has been reported that human breast milk
contains a number of oligosaccharides that are only digested by gut bacteria,
not the breast-feeding child [19,20]. Therefore, recent publications that
report gluten-degrading bacteria as natural residents of the oral cavity are
not surprising after all [21,22]. This discovery is also very significant, since
the oral cavity represents the gateway to the gastro-intestinal system in which
gluten is mixed with the oral
microorganisms in human saliva. The finding of gluten-degrading oral
microbes then begs the questions, what are they susceptible to and what common
source may reduce the gluten metabolizers or decrease their effectiveness of
gluten processing, leading to gluten “sensitivity”? Susceptibility
Experiment Three colonies of R. aeria,
R. dentocariosa, R. mucilaginosa, S. mutans,
or Lactobacillus were obtained from
isolation plates and grown in Mueller-Hinton media to a McFarland Standard of
0.5. Either Brucella
agar plates, Rogosa agar, or Mueller-Hinton agar plates with 5% sheep blood
were wholly spread to create a lawn with one cotton swab inoculation of chosen
target bacteria. Five cotton discs were evenly distributed on the plate and 10
microliters of full strength OTC reagent was pipetted directly onto each
corresponding disc. The plates were evaluated after 30 hours of growth at 36oC.
Calipers were used to measure zones of inhibition in millimeters. Diffusion
Experiment Trypticase Soy Agar (TSA) was autoclaved and cooled to 56
degrees and aliquots of 25 mL were cooled and inoculated with 2 mL of 0.5
McFarland Standard suspensions of target organisms: R. dentocariosa, R. mucilaginosa,
Streptococcus salivarius, Escherichia coli or Pseudomonas
aeruginosa prior to pouring agar plates. Impregnated plates were then
inoculated in punched zones using a disposable 10 microliter loop with 0.5
McFarland Standards of test inhibiting bacteria species: Streptococcus salivarius, Staphylococcus
aureus, Vancomycin-resistant Enterococcus, Pseudomonas aeruginosa, Escherichia
coli, and R. dentocariosa or R. mucilaginosa. The plates were
evaluated after 24 hours of growth at 36oC. Calipers were used to
measure zones of inhibition. Spry™ Xylitol Toothpaste Gel inhibited R. mucilaginosa. L. reuteri,
a probiotic that inhibits many oral and pathogens, was significantly inhibited
by OTC oral products, except the xylitol based. Xylitol based oral products did
not inhibit the commensal S.
salivarius nor the pathogens, S.
aureus, E. coli and P. aeruginosa (Table 2). Xylitol based oral products do inhibit many oral pathogens
and have been extensively used in dentistry for decades. Growth of P.
aeruginosa was inhibited by R. dentocariosa
and growth of S. aureus was inhibited
by R. mucilaginosa. The zones of
inhibition by the gluten metabolizers were demonstrably large. The inhibition
of the beneficial gluten metabolizers and probiotic bacteria by OTC oral
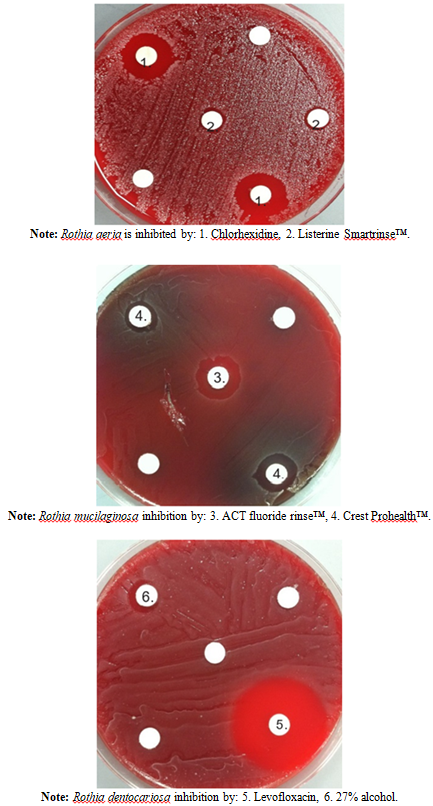
products may have been the result of fluoride concentration. An alcohol based product, ListerineTM, did not
greatly inhibit the gluten metabolizers (Figure
1). Figure 1: Example of Inhibition of pathogen P. aeruginosa by gluten metabolizer, R. dentocariosa. In vitro results are not always applicable
to the clinical situation. Indeed, the complexity of the human oral microbiome
would make it difficult to predict a response to any oral intervention with
certainty. The results of the present study are of a pilot nature, a negative
finding would mean that there is little need for further investigation.
However, in vitro studies are always
necessary before progressing into more extensive, time consuming, and
financially demanding clinical studies. The mere fact that OTC products,
sometimes used ad libitum by patients, contribute to a reduction in beneficial
bacteria should be a concern to all health practitioners. Of greater interest
should be the extent of inhibition, as the zones of inhibition were quite
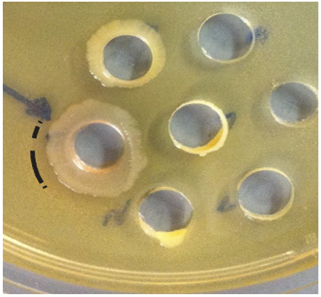
significant in diameter. The average
diameter of inhibition with an OTC product was 13 mm [Range: <6-18 mm] (Figure 2). Figure 2:Examples of Inhibition Plates by
OTC Products. The mode of inhibition should be discovered, as it appears
that the fluoride concentration of the OTC products may have been contributory.
An alcohol based product was only inhibitory of the probiotic in this study,
and not the gluten metabolizers. With dental disease at an increasing rate in
developing countries due to the shift to a higher carbohydrate diet, with
addition of processed foods containing added sugars, health professionals
should be cautioning about the over use of OTC
products. The dental caries rate is not decreasing, as would be expected
with all the OTC utilization, and dental expenditures are increasing every year. Perhaps the OTC products help with
limiting the pathogenic bacteria but only at the expense of also eliminating
many beneficial bacteria. This is a no win situation for the population,
spending vital resources on products that may indeed create more pathology,
such as, gluten sensitivity, and fail to protect from dental caries. The beneficial effect of fluoride for caries protection may
be somewhat decreased by the possible inhibition of oral probiotic bacteria by
over use of OTC products. Unsupervised use of a daily fluoride mouth rinse by a
child could possibly create a gluten sensitivity issue, and due to lack of
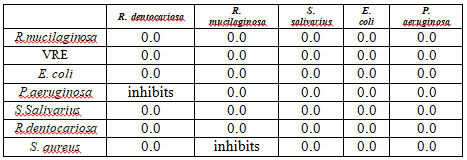
regulatory oversight, this severe side effect would never be discovered (Table 3). Table 3:Diffusion experiment: Bacterial
species inhibition of each other. Another very important aspect of this study was the
interaction between pathogenic and beneficial bacteria. The interaction, or
rather, the inhibition of different bacterial species actually determines the
health of the host and as such, is paramount in importance. The results were
significant in that growth of Rothia
species was inhibited by other bacteria. This suggests that if the oral flora
equilibrium is changed by using OTC oral hygiene products, a domino effect can
change the entire oral microbiome, which is the gateway to the digestive tract.
The gastric microbiome is now recognized as a vital component of the host’s
health, both mental and physical. Increased oversight concerning the over uses of
anti-microbial, food
preservatives that are also anti-microbial, and OTC products that inhibit
commensal bacteria, is essential. Required testing of OTC products and better
population education into the importance of the holobiome should be a health
priority. The connection between the increase in chronic diseases and the
significant shift reported in the modern human microbiome should be further
investigated. Gluten oral bacteria, Rothia mucilaginosa and
Streptococcus salivarius.Inhibition of Rothia Species by Over-the-Counter Products and Bacterial Antagonists
Abstract
Full-Text
Introduction
Objective
The purpose of this study was to determine if there is any
inhibition of beneficial oral biofilm species such as Rothia aeria, R. mucilaginosa and R. dentocariosa, Streptococcus
mutans (pathogen-negative control) and also Lactobacillus reuteri strains (isolated from periobalance
Probiotic) by Over The Counter (OTC) oral antimicrobials
utilizing in vitro laboratory
technique. The secondary objective was to determine the antagonism, if any, of
the Rothia genus by Streptococcus species (mutans and salivarius) and known pathogens.
Rothia aeria and R. mucilaginosa are reported to be
important in the processing of gluten. Inhibition of these beneficial bacteria
by OTC products, either directly or indirectly, would increase gluten
sensitivity in patients. Beneficial bacteria may be indirectly inhibited by
certain antagonistic bacteria that are relatively less sensitive to OTC
products.
Methods
Results
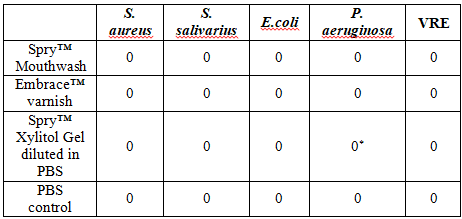
Bacterial growths of all tested bacteria were inhibited by
Crest ProHealth™, ACT™, Listerine SmartRinse™, and Chlorhexidine. R. aeria and R. mucilaginosa were also inhibited by Embrace™ varnish (Table 1).

Discussion
Conclusion
Rothia and Lactobacillus species may be decreased in quantity by the overuse
of oral antimicrobials. OTC products may alter the oral microbiome creating a
situation less conducive for the survival of essential beneficial bacteria. The
use of OTC products may decrease the enzymatic degradation of gluten containing
foods by Rothia bacteria. This can
possibly result in gluten sensitivity, thereby increasing the clinical
prevalence of celiac disease. Further studies are required before any clinical
implications may be concluded, but oral antimicrobials should be used only when
necessary.
References
*Corresponding author
Mark L Cannon,
Professor, Feinberg School of Medicine, Northwestern University, Chicago, USA,
Tel: 847-899-6720, E-mail: drmarkcannon@outlook.com
Citation
Mark LC, Kabat B, Yogev R, Jantra L, Awan A, et al. Inhibition of
rothia species by over-the-counter products and bacterial antagonists (2020) Edelweiss Appli Sci4: 5-8. Keywords